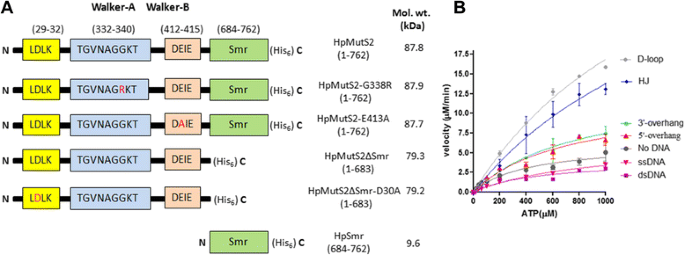
Mutations in the nucleotide binding and hydrolysis domains of Helicobacter pylori MutS2 lead to altered biochemical activities and inactivation of its in vivo function | BMC Microbiology | Full Text
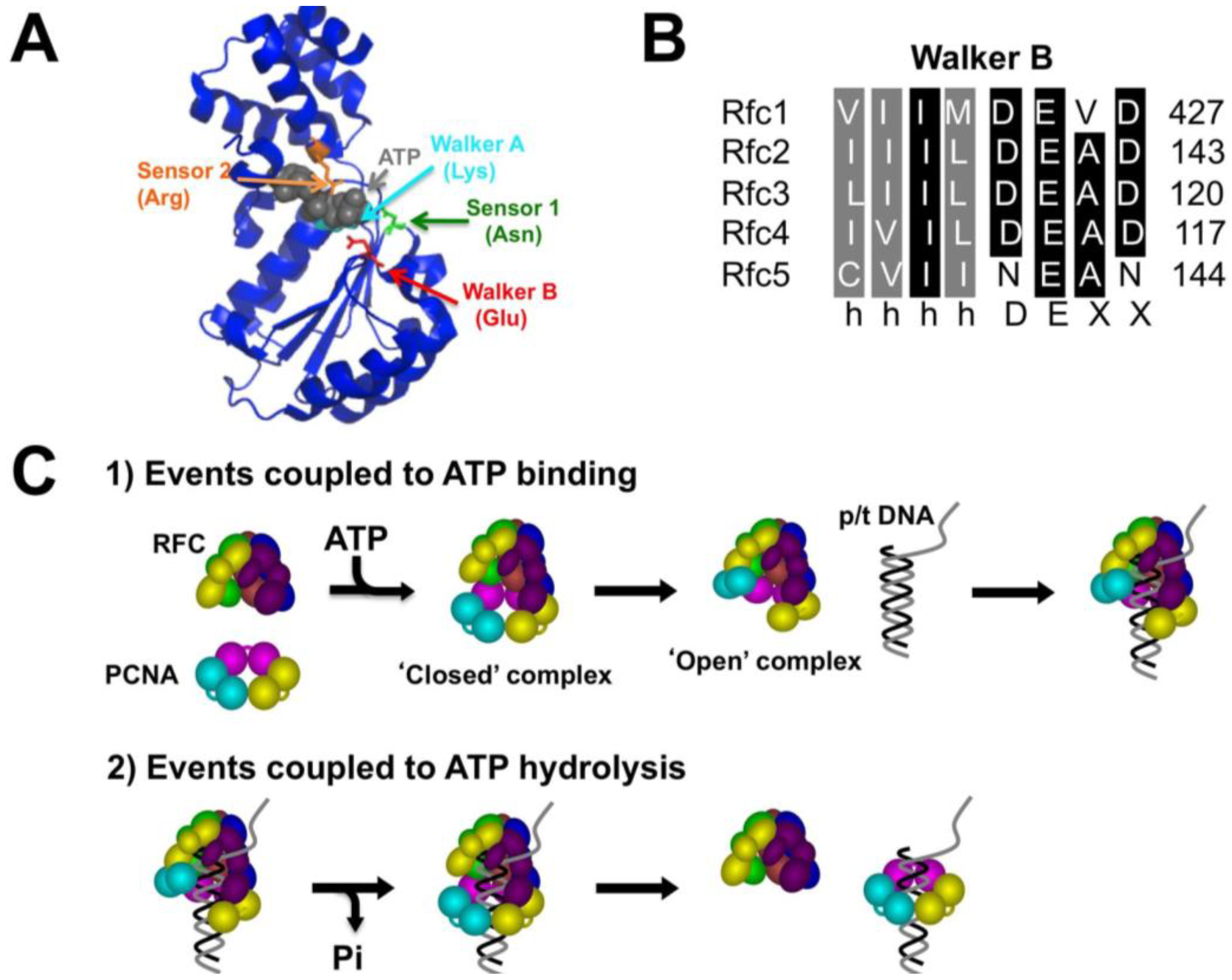
Genes | Free Full-Text | A Novel Function for the Conserved Glutamate Residue in the Walker B Motif of Replication Factor C
The ATPase site and mutations affecting SecA function. (a) The Walker A... | Download Scientific Diagram
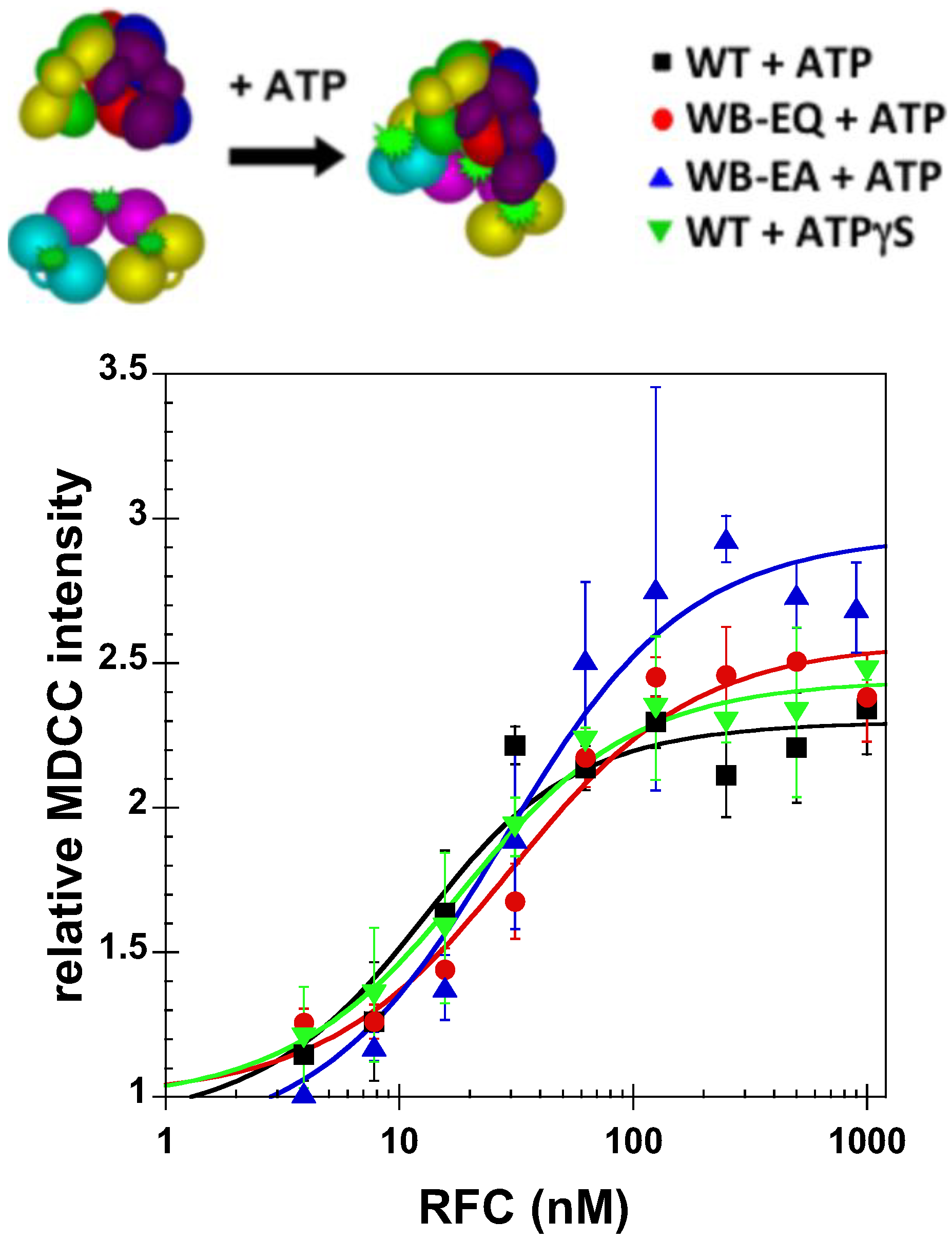
Genes | Free Full-Text | A Novel Function for the Conserved Glutamate Residue in the Walker B Motif of Replication Factor C

Toward wide-spectrum antivirals against coronaviruses: Molecular characterization of SARS-CoV-2 NSP13 helicase inhibitors | Science Advances

The Conserved Glutamate Residue Adjacent to the Walker-B Motif Is the Catalytic Base for ATP Hydrolysis in the ATP-binding Cassette Transporter BmrA* - Journal of Biological Chemistry

Model of the NOD2 nucleotide-binding site with an ADP molecule and conserved sequence motifs Walker A, Walker B, Sensor 1, GxP, and WH-His shown in sticks.
![PDF] The Walker B motif of the second nucleotide-binding domain (NBD2) of CFTR plays a key role in ATPase activity by the NBD1-NBD2 heterodimer. | Semantic Scholar PDF] The Walker B motif of the second nucleotide-binding domain (NBD2) of CFTR plays a key role in ATPase activity by the NBD1-NBD2 heterodimer. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/fb03c57d46cf8f1d0e5151dfa451ac68d4a2966f/3-Figure1-1.png)
PDF] The Walker B motif of the second nucleotide-binding domain (NBD2) of CFTR plays a key role in ATPase activity by the NBD1-NBD2 heterodimer. | Semantic Scholar
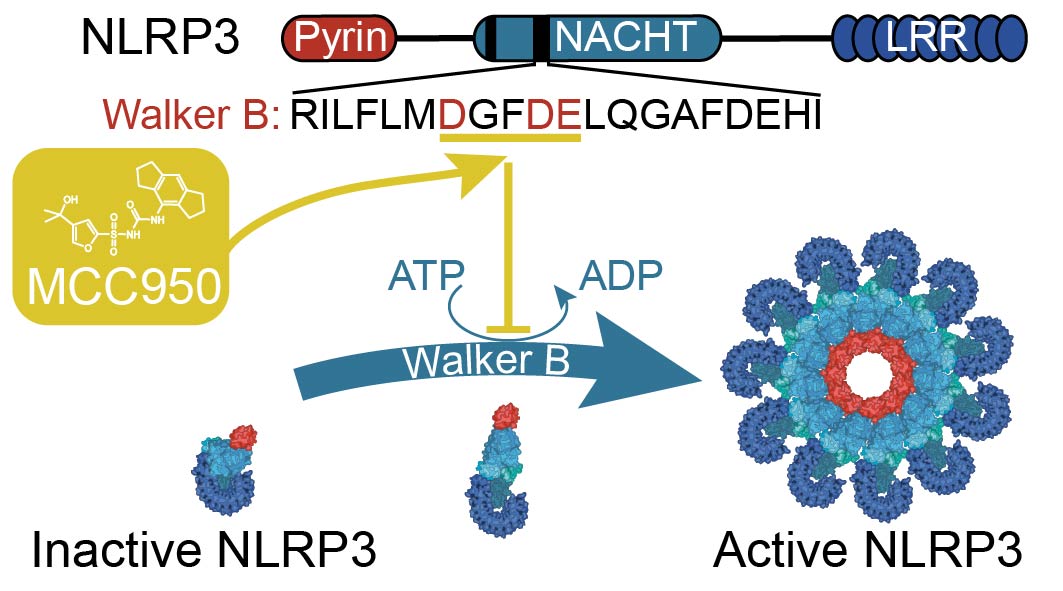
MCC950 directly targets the NLRP3 ATP-hydrolysis motif for inflammasome inhibition | IMB Inflammasome Lab

Walker-A Motif Acts to Coordinate ATP Hydrolysis with Motor Output in Viral DNA Packaging - ScienceDirect

The putative Walker A and Walker B motifs of Rrp2 are required for the growth of Borrelia burgdorferi - Ouyang - 2017 - Molecular Microbiology - Wiley Online Library

Visualization of NBD motifs: The NBD-NBD interface of a single NBD is shown in cartoon representation.

A Distinct Motif in a Prokaryotic Small Ras-Like GTPase Highlights Unifying Features of Walker B Motifs in P-Loop NTPases - ScienceDirect

Figure 1 from Role of highly central residues of P-loop and it's flanking region in preserving the archetypal conformation of Walker A motif of diverse P-loop NTPases | Semantic Scholar

Understanding Protein Function - The Disparity Between Bioinformatics and Molecular Methods | IntechOpen

A Distinct Motif in a Prokaryotic Small Ras-Like GTPase Highlights Unifying Features of Walker B Motifs in P-Loop NTPases - ScienceDirect